OAH & 3DM
Index
General
For this exercise, you need either Pymol or Yasara installed that has the 3DM plugin. If you don't have Yasara or Pymol or you are missing the 3DM functionality, please consult the installation instructions. Before you start this exercise make sure you have the latest version of Yasara or Pymol installed.
Login at 3DM with your 3DM account. If you don't have a 3DM account you can request one via the "get 3DM" tab. To be able to do this course you need to have access to the databases used in this course. After you have requested an account you can request access to the course databases by sending an email to joosten@bio-prodict.nl.
After entering the login details you will see a 'Select 3DM system' page. Click on the 'Public 3DM Systems' checkbox and search for the "Phosphoenolpyruvate mutase/Isocitrate lyase" database. Open this 3DM system.
At the starting page of each 3DM database, you see the 3DM data cycle. The icons in the circle represent links to the most important 3DM options. These options are also available on the left.
Introduction
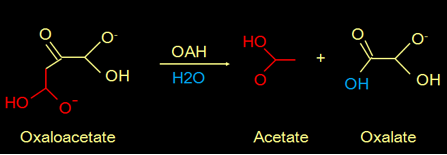
Fungi can be pathogenic to plants and animals. It is known that the secretion of oxalate by fungi is a commonly used strategy for their pathogenicity. Oxalate is toxic and can form crystals that demolish the cell wall of the host. The oxalate is produced from oxaloacetate catalyzed by the enzyme oxaloacetate hydrolase (OAH). This is the reaction:
Fig 1. Reaction mechanism that produces oxalate.
We have generated a 3DM for the corresponding protein family. OAH falls in the Phosphoenolpyruvate mutase/Isocitrate lyase superfamily.
The OAH of niger is the best characterized OAH protein. This is the sequence:
>G3Y473 MKVDTPDSASTISMTNTITITVEQDGIYEINGARQEPVVNLNMVTGASKLRKQLRETNEL LVCPGVYDGLSARIAINLGFKGMYMTGAGTTASRLGMADLGLAHIYDMKTNAEMIANLDP YGPPLIADMDTGYGGPLMVARSVQQYIQAGVAGFHIEDQIQNKRCGHLAGKRVVTMDEYL TRIRAAKLTKDRLRSDIVLIARTDALQQHGYDECIRRLKAARDLGADVGLLEGFTSKEMA RRCVQDLAPWPLLLNMVENGAGPVISVDEAREMGFRIMIFSFACITPAYMGITAALERLK KDGVVGLPEGMGPKKLFEVCGLMDSVRVDTEAGGDGFANGV
For each protein in the 3DM database, there is a "protein information" page that contains more detailed information.
On the protein information pages, you can find a couple of different tabs. Have a quick look at what you can find in each tab.
Subsets
3DM offers several ways to select a subset of sequences. Once a subset is selected a mini 3DM can be generated for this subset. All 3DM functionalities, such as the correlated mutations, are regenerated and can separately be analyzed. The data of a subset can also be compared to the data of the full set of sequences or with other previously defined subsets.
With the search option we have made a subset called "oxalate producers" that contains the proteins available in this 3DM system for fungi of which it is known that they can produce oxalate:
Aspergillus clavatus Neosartorya fischeri Penicillium chrysogenum Penicillium marneffei Talaromyces stipitatus Sclerotinia sclerotiorum Aspergillus niger Sclerotium cepivorum Aspergillus terreus Aspergillus fumigatus Botryotinia fuckeliana
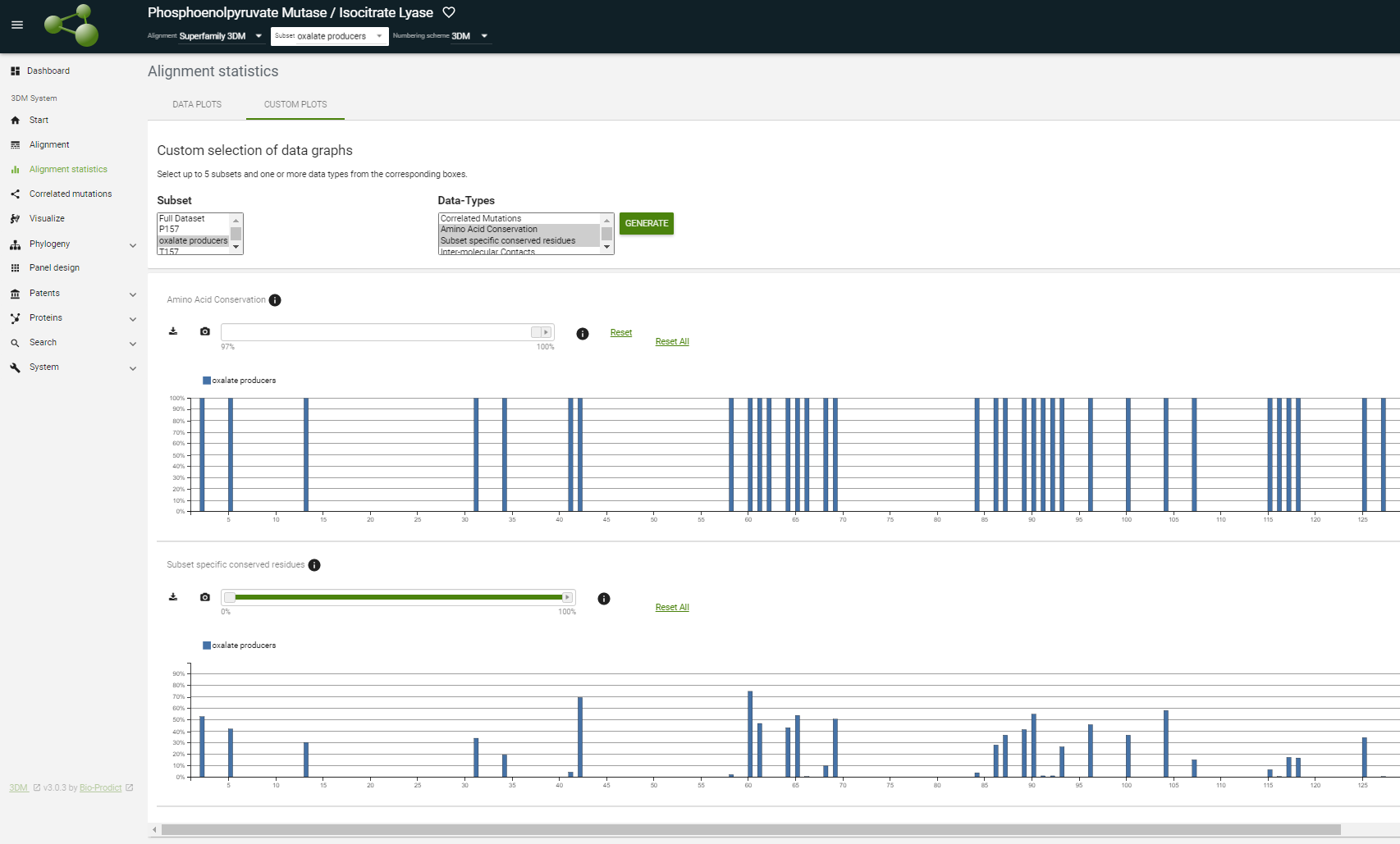
At the alignment, statistics pages change between the full dataset and the "oxalate producers" you just made using the 'Subset' menu in the header on top of 3DM and see how the graphs change.
3DM always generates an extra histogram for each subset that shows the residues that are specifically conserved in the selected subset (the histogram called "subset specific conserved residues"). The highest scoring residues are around 3D positions 157.
Important here is to realize that these are positions that are not just simply conserved in this subset of oxalate producing fungi, but the corresponding residues are absent from the rest of the sequences in the superfamily. In other words, these residues are specific for this subset.
You can see this by comparing this plot with the amino acid conservation plot of the new subset. Use the "custom plot" tab and select your subset from the left box and from the right box "amino acid conservation " and "subset specific conserved residues".
On the alignment page:
- Click on the consensus sequence at position 157
Take home message: the data you are looking at is always depending on the subset tab that is selected.
Click on "Correlated mutations" in the menu on the left. Make sure you have the "Full Dataset" selected at the top of 3DM.
Correlated mutations calculated for a superfamily alignment often reflect positions important for specificity because superfamily alignments contain enzymes with different specificities (do you understand this concept?)
The "Top Correlation Heatmap" page shows the alignment positions of which the residues mutate simultaneously (definition of a correlated mutation).
- Select the "Correlation Networks" tab. Here you can give a keyword in the "Literature & Mutations" window on the right.
- Type " specificity" in the box. This will select mutations from the literature that affect specificity reported in any of the proteins of the superfamily.
- Go back to the alignment statistics page.
- Now compare the ligand contact plot (in this case these will be enzyme inhibitors) of the full dataset with the correlation plot of the full dataset
Homology modeling of OAH niger with its substrate oxaloacetate and the design of an inhibitor
Build a homology model
- Go back to the protein information page of G3Y473 and select the ‘MODELS' tab.
3DM selected three structures as potential good templates. In this course, you will learn how to select the best template, make the best alignments, etc., but for now, we will use 3LYEA → it does have the best resolution (e.g. quality).
Select 3LYEA as a template and use the "alignment" as numbering. You can open the model either with Yasara or Pymol. Note that generated models can always be retrieved from the "visualize" pages. The third form on this page contains the models. Here you can select your model and different data that you want to visualize in your model.
- Select in yasara or pymol residue with 3D number 157.
Load an inhibitor
Structures can be loaded directly in Yasara from the 3DM database via the 3DM → Structures → load structure from 3DM option. Loading structure files via the 3DM menu ensures that the structures are all superimposed, co-crystallized compounds will have to be positioned in the active site and proteins will have the 3D numbering.
- Load the inhibitor of 1M1BA using the "load data from 3DM" option in Yasara.
Build oxaloacetate from this inhibitor.
Fig 2. Structure of oxaloacetate.
This is the structure of oxaloacetate. We are very lucky since it is very similar to the structure of the 1M1BA inhibitor. Simply swapping the SO3 group with a CO2 group will do the job.
In Yasara:
- To do this delete one oxygen of SO3 → select it and press delete.
- Then right-click on the S and select "swap → atom" and replace it with carbon. The angles are not perfect (it needs energy minimization), but it gives a quick and dirty idea how oxaloacetate fits in the active site.
In Pymol:
- Load the 1M1B structure
- Zoom in on the ligand, find the SO3 group
- Ctrl + Middle click on one of the Oxygen atoms on the SO3 group. A number of extra objects appear in the object list on the right.
- In the command line at the top, enter: remove pk1 and press enter. The oxygen atom will disappear.
- Ctrl + Middle-click on the S atom in the group.
- In the command line, enter: alter pk1,elem="C" , then press Enter
- In the command line, enter: alter pk1,name="C4" , then press Enter
- In the object list, click on the C that appears next to the 1M1B object. Select any of the coloring schemes under Color... By element. The SO3 group will now be colored the same as a CO2 group.
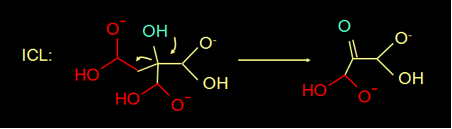
The reaction mechanism of isocitrate lyase (ICL) is known for quite a while (fig 3). In this reaction mechanism the H of the blue OH group donates an electron, makes a double bond, and splits of the COOH group.
Fig 3. Reaction mechanism of ICL (above) and the structure of oxaloacetate (below)
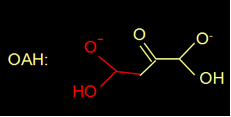
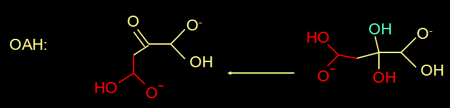
Actually, oxaloacetate in water is in equilibrium with its diol form (figure 4).
Fig 4. Oxaloacetate is in equilibrium with its diol.
Until today OAH is the only known enzyme of this superfamily that has a substrate in a diol form. So the extra OH is unique to OAH.
Modeling the extra OH in the active site with the "swap" option does not work very well in yasara, because yasara can't deal with changing the double bond of C=O to the single bond of C-OH without proper energy minimization (try to make the diol with the swap option if you like).
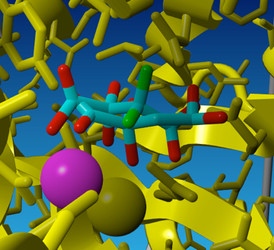
Fig 5. The result of energy minimization performed on the diol form of oxaloacetate in the OAH model.
In 2008 a model of OAH was generated similar to the way you did it today. With this model we were already in 2008 able to:
- Reveal the OAH specific serine 157 (figure 6)
- Reveal the reaction mechanism of OAH (via the diol substrate)
- Show the relation between oxalate production and pathogenicity of fungi
- Make a very strong inhibitor of OAH (potential anti-fungal drug)
The inhibitor was designed by organic chemists that realized they had to make a compound that is 100% in the diol form. This was the case with difluoro-oxaloaceate. This compound indeed proved to be a very strong inhibitor of OAH and was later crystallized together with OAH of the fungus Cryphonectria Parasitica (pdb file 3M0JA).
- To see how well you modeled oxaloacetate in the active site load the drug of 3M0JA in your model with the 3DM option of Yasara.
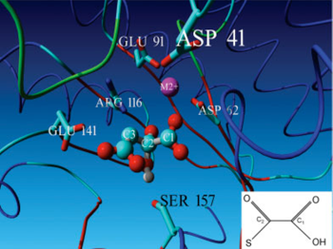
Fig 6. Picture of the model of OAH taken from the 2008 publication: Identification of fungal oxaloacetate hydrolyase within the isocitrate lyase/PEP mutase enzyme superfamily using a sequence marker-based method. This picture clearly shows the predicted Ser157 H-bridge with the diol of oxaloacetate.
Extra questions
Position 157 is the center of the correlated mutation network. P is the most common residue at position 157 (is that correct?). We have generated a subset of sequences that have a P at position 157 called "P157"
Take home message: The function underlying correlated mutations heavily depend on the input alignment. Always look for additional data (in this case protein-protein interaction data → did you find that?) that might explain a correlated mutation network.
The correlated mutations in this superfamily seem to reflect positions important for specificity. You want to change the specificity of OAH and you decide to rationally design a mutant library. Your screening method allows you to screen up to 1000 mutant clones.